| E2F7 |
|---|
|
| Identifiers |
|---|
| Aliases | E2F7, E2F transcription factor 7 |
|---|
| External IDs | OMIM: 612046; MGI: 1289147; HomoloGene: 18685; GeneCards: E2F7; OMA:E2F7 - orthologs |
|---|
| Gene location (Human) |
|---|
 | | Chr. | Chromosome 12 (human)[1] |
|---|
| | Band | 12q21.2 | Start | 77,021,248 bp[1] |
|---|
| End | 77,065,569 bp[1] |
|---|
|
| Gene location (Mouse) |
|---|
 | | Chr. | Chromosome 10 (mouse)[2] |
|---|
| | Band | 10 D1|10 57.74 cM | Start | 110,581,300 bp[2] |
|---|
| End | 110,623,245 bp[2] |
|---|
|
| RNA expression pattern |
|---|
| Bgee | | Human | Mouse (ortholog) |
|---|
| Top expressed in | - ventricular zone
- thymus
- gonad
- ganglionic eminence
- testicle
- secondary oocyte
- stromal cell of endometrium
- mucosa of ileum
- bone marrow cells
- pancreatic ductal cell
|
| | Top expressed in | - otic vesicle
- hand
- primitive streak
- fossa
- tail of embryo
- condyle
- yolk sac
- genital tubercle
- abdominal wall
- mandibular prominence
|
| | More reference expression data |
|
|---|
| BioGPS | 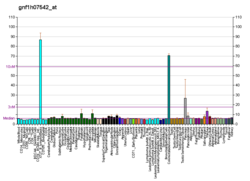 | | More reference expression data |
|
|---|
|
| Gene ontology |
|---|
| Molecular function | - DNA binding
- RNA polymerase II transcription regulatory region sequence-specific DNA binding
- transcription corepressor activity
- protein binding
- DNA-binding transcription repressor activity, RNA polymerase II-specific
- identical protein binding
- DNA-binding transcription factor activity
- DNA-binding transcription factor activity, RNA polymerase II-specific
- RNA polymerase II cis-regulatory region sequence-specific DNA binding
- cis-regulatory region sequence-specific DNA binding
- DNA-binding transcription activator activity, RNA polymerase II-specific
- transcription factor binding
- sequence-specific DNA binding
| | Cellular component | - transcription regulator complex
- nucleus
- nucleoplasm
- nuclear speck
- RNA polymerase II transcription regulator complex
| | Biological process | - trophoblast giant cell differentiation
- regulation of transcription, DNA-templated
- chorionic trophoblast cell differentiation
- negative regulation of transcription involved in G1/S transition of mitotic cell cycle
- placenta development
- DNA damage response, signal transduction by p53 class mediator
- hepatocyte differentiation
- negative regulation of transcription by RNA polymerase II
- transcription, DNA-templated
- cellular response to DNA damage stimulus
- negative regulation of G1/S transition of mitotic cell cycle
- cell cycle
- negative regulation of cytokinesis
- positive regulation of DNA endoreduplication
- positive regulation of transcription by RNA polymerase II
- sprouting angiogenesis
- DNA damage response, signal transduction by p53 class mediator resulting in cell cycle arrest
- negative regulation of cell population proliferation
- regulation of cell cycle
| | Sources:Amigo / QuickGO |
|
| Orthologs |
|---|
| Species | Human | Mouse |
|---|
| Entrez | | |
|---|
| Ensembl | | |
|---|
| UniProt | | |
|---|
| RefSeq (mRNA) | | |
|---|
| RefSeq (protein) | | |
|---|
| Location (UCSC) | Chr 12: 77.02 – 77.07 Mb | Chr 10: 110.58 – 110.62 Mb |
|---|
| PubMed search | [3] | [4] |
|---|
|
| Wikidata |
| View/Edit Human | View/Edit Mouse |
|