| TLE4 |
|---|
 |
| Identifiers |
|---|
| Aliases | TLE4, BCE-1, BCE1, E(spI), ESG, ESG4, GRG4, Grg-4, transducin like enhancer of split 4, TLE family member 4, transcriptional corepressor, E(spl) |
|---|
| External IDs | OMIM: 605132; MGI: 104633; HomoloGene: 38259; GeneCards: TLE4; OMA:TLE4 - orthologs |
|---|
| Gene location (Human) |
|---|
 | | Chr. | Chromosome 9 (human)[1] |
|---|
| | Band | 9q21.31 | Start | 79,571,773 bp[1] |
|---|
| End | 79,726,882 bp[1] |
|---|
|
| Gene location (Mouse) |
|---|
 | | Chr. | Chromosome 19 (mouse)[2] |
|---|
| | Band | 19 A|19 9.11 cM | Start | 14,425,514 bp[2] |
|---|
| End | 14,575,415 bp[2] |
|---|
|
| RNA expression pattern |
|---|
| Bgee | | Human | Mouse (ortholog) |
|---|
| Top expressed in | - optic nerve
- left testis
- right testis
- endothelial cell
- corpus callosum
- inferior olivary nucleus
- inferior ganglion of vagus nerve
- dorsal motor nucleus of vagus nerve
- Region I of hippocampus proper
- bone marrow cells
|
| | Top expressed in | - renal corpuscle
- medullary collecting duct
- vestibular membrane of cochlear duct
- superior cervical ganglion
- suprachiasmatic nucleus
- conjunctival fornix
- nucleus accumbens
- retinal pigment epithelium
- tail of embryo
- olfactory tubercle
|
| | More reference expression data |
|
|---|
| BioGPS | 
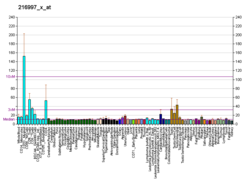 | | More reference expression data |
|
|---|
|
| Gene ontology |
|---|
| Molecular function | - transcription factor activity, RNA polymerase II distal enhancer sequence-specific binding
- chromatin binding
- protein binding
- transcription corepressor activity
- DNA-binding transcription factor activity, RNA polymerase II-specific
- molecular function
| | Cellular component | - nucleus
- nucleoplasm
- transcription regulator complex
- beta-catenin-TCF complex
| | Biological process | - negative regulation of transcription, DNA-templated
- regulation of transcription, DNA-templated
- negative regulation of transcription by RNA polymerase II
- Wnt signaling pathway
- transcription, DNA-templated
- beta-catenin-TCF complex assembly
- cellular response to leukemia inhibitory factor
- biological process
- negative regulation of canonical Wnt signaling pathway
| | Sources:Amigo / QuickGO |
|
| Orthologs |
|---|
| Species | Human | Mouse |
|---|
| Entrez | | |
|---|
| Ensembl | | |
|---|
| UniProt | | |
|---|
| RefSeq (mRNA) | NM_001282748
NM_001282749
NM_001282753
NM_001282760
NM_007005
|
|---|
NM_001351541
NM_001351542
NM_001351543
NM_001351546
NM_001351547
NM_001351550
NM_001351552
NM_001351558
NM_001351560
NM_001351562
NM_001351563
NM_001351564
NM_001351556 |
| |
|---|
NM_011600
NM_001302947
NM_001302950
NM_001302951 |
|
|---|
| RefSeq (protein) | NP_001269677
NP_001269678
NP_001269682
NP_001269689
NP_008936
|
|---|
NP_001338470
NP_001338471
NP_001338472
NP_001338475
NP_001338476
NP_001338479
NP_001338481
NP_001338487
NP_001338489
NP_001338491
NP_001338492
NP_001338493
NP_001338485 |
| |
|---|
NP_001289876
NP_001289879
NP_001289880
NP_035730 |
|
|---|
| Location (UCSC) | Chr 9: 79.57 – 79.73 Mb | Chr 19: 14.43 – 14.58 Mb |
|---|
| PubMed search | [3] | [4] |
|---|
|
| Wikidata |
| View/Edit Human | View/Edit Mouse |
|

 1gxr: WD40 REGION OF HUMAN GROUCHO/TLE1
1gxr: WD40 REGION OF HUMAN GROUCHO/TLE1 2ce8: AN EH1 PEPTIDE BOUND TO THE GROUCHO-TLE WD40 DOMAIN.
2ce8: AN EH1 PEPTIDE BOUND TO THE GROUCHO-TLE WD40 DOMAIN. 2ce9: A WRPW PEPTIDE BOUND TO THE GROUCHO-TLE WD40 DOMAIN.
2ce9: A WRPW PEPTIDE BOUND TO THE GROUCHO-TLE WD40 DOMAIN.




















